| Version 4 (modified by , 7 weeks ago) ( diff ) |
|---|
Adapting a model's parallelization for PDAF
Online Mode: Implementation Guide
- Main page
- Adapting the parallelization
- Initializing PDAF
- Modifications for ensemble integration
- Implementing the analysis step
- Memory and timing information
Contents of this page
| This page described the adaption of the model parallelization for PDAF V3.0 and later. For PDAF V2.x, see the Page on adapting the parallelization in PDAF 2 |
Overview
The PDAF release provides example code for the online mode in tutorial/online_2D_serialmodel (for a model without parallelization) and tutorial/online_2D_parallelmodel (for a parallelized model). We refer to this code to use it as a basis.
In the tutorial code and the templates in templates/online, the parallelization is initialized in the routine init_parallel_pdaf (file init_parallel_pdaf.F90). The required variables are defined in mod_parallel_pdaf.F90. These files can be used as templates.
In implementations done with PDAF V3.1 and later init_parallel_pdaf calls the routine PDAF3_init_parallel to configure the parallelization for PDAF. In PDAF V3.0 and before the init_parallel_pdaf contains the actual configuration of the parallelization. In PDAF V3.0 the parallelization variables are then provided to PDAF by a call to PDAF3_set_parallel. Implementations done before do usually not include this call, but provide the variables to PDAF in the initialization call to PDAF_init.
Like many numerical models, PDAF uses the MPI standard for the parallelization. For the case of a parallelized model, we assume in the description below that the model is parallelized using MPI. If the model is parallelized using OpenMP, one can follow the explanations for a non-parallel model below.
As explained on the page on the Implementation concept of the online mode, PDAF supports a 2-level parallelization. First, the numerical model can be parallelized and can be executed using several processors. Second, several model tasks can be computed in parallel, i.e. a parallel ensemble integration can be performed. We need to configure the parallelization so that more than one model task can be computed.
There are two possible cases regarding the parallelization for enabling the 2-level parallelization:
- The model itself is parallelized using MPI
- In this case we need to adapt the parallelization of the model for the data assimilation with PDAF
- The model is not parallelized, i.e. a serial model, or uses only shared-memory parallelization using OpenMP
- In this case we use PDAF to add parallelization
Adaptions for a parallelized model
If the online mode is implemented with a parallelized model, one has to ensure that the parallelization can be split to perform the parallel ensemble forecast. For this, one has to check the model source code and potentially adapt it.
The adaptions in short
If you are experienced with MPI, the steps are the following:
- Check whether the model uses
MPI_COMM_WORLD.- If yes, then replace
MPI_COMM_WORLDin all places, exceptMPI_abortandMPI_finalizeby a user-defined communicator (we call itCOMM_mymodelhere), which can be initialized asCOMM_mymodel=MPI_COMM_WORLD. - If no, then take note of the name of the communicator variable (we assume here it's
COMM_mymodel).
- If yes, then replace
- Find the call to
MPI_init - Insert the call
init_parallel_pdafdirectly afterMPI_initand possible calls toMPI_Comm_sizeandMPI_Comm_rankprovidingCOMM_mymodeland the related rank and size variables. - The number of model tasks in variable
n_modeltasksis required byPDAF3_init_parallelto perform communicator splitting. In the tutorial code we include a command-line parsing to set the variable (parsed with the keyworddim_ens, thus specifying-dim_ens N_MODELTASKSon the command line, where N_MODELTASKS should be the ensemble size). One could also read the value from a configuration file.
Details on adapting a parallelized model
Any program parallelized with MPI will need to call MPI_Init for the initialziation of MPI.
Frequently, the parallelization of a model is initialized in the model by the lines:
CALL MPI_Init(ierr)
CALL MPI_Comm_rank(MPI_COMM_WORLD, rank, ierr)
CALL MPI_Comm_size(MPI_COMM_WORLD, size, ierr)
Here, the call to MPI_init is mandatory, while the two other lines are optional, but common. The call to MPI_init initializes the parallel region of an MPI-parallel program. This call initializes the communicator MPI_COMM_WORLD, which is pre-defined by MPI to contain all processes of the MPI-parallel program.
In the model code, we have to find the place where MPI_init is called to check how the parallelization is set up. In particular, we have to check if the parallelization is ready to be split into model tasks.
For this one has to check if MPI_COMM_WORLD, e.g. checking the calls to MPI_Comm_rank and MPI_Comm_size, or MPI communication calls in the code (e.g. MPI_Send, MPI_Recv, MPI_Barrier).
- If the model uses
MPI_COMM_WORLDwe have to replace this by a user-defined communicator, e.gCOMM_mymodel. One has to declare this as an integer variable in a module and initialize it asCOMM_mymodel=MPI_COMM_WORLD. Then all occurences ofMPI_COMM_WORLDin the code are replaced byCOMM_mymodel. This change must not influence the execution of the model, as on e can check with a test run. - If the model uses a different communicator, one should take note of its name (below we refer to it as
COMM_mymodel). PDAF will reconfigure it to be able to represent the ensemble of model tasks.
Now, we add the call to init_parallel_pdaf which reconfigures COMM_mymodel and the related rank and size variables. These variables are arguments to the template variant of init_parallel_pdaf (i.e `template/online/init_parallel_pdaf.F90)
Finally, we have a ensure that the number of model tasks is correctly set. In the template and tutorial codes, the number of model tasks is specified by the variable n_modeltasks. This has to be set before the operations on communicators are done in init_parallel_pdaf. In the tutorial code we added a command-line parsing to set the variable (parsed with the keyword dim_ens, thus specifying -dim_ens N_MODELTASKS on the command line, where N_MODELTASKS should be the ensemble size). One could also read the value from a configuration file.
If the program is executed with these extensions using multiple model tasks, the issues discussed in 'Compiling the extended program' can occur. This one has to take care about which processes will perform output to the screen or to files.
Adaptions for a serial model
The adaptions in short
If you are experienced with MPI, the steps are the following using the files mod_parallel_pdaf.F90, init_parallel_pdaf.F90.serialmodel, finalize_pdaf.F90 and parser_mpi.F90 from tutorial/online_2D_serialmodel.
- rename the file
init_parallel_pdaf.F90.serialmodeltoinit_parallel_pdaf.F90when copying it. - Insert
CALL init_parallel_pdaf(1)at the beginning of the main program. This routine performs the initialization of MPI and the configuration of communicators for the data assimilation - Insert
CALL finalize_pdaf()at the end of the main program. This routine will also finalize MPI. - The number of model tasks in variable
n_modeltasksis determined ininit_parallel_pdafby command line parsing (it is parsing fordim_ens, thus setting-dim_ens N_MODELTASKSon the command line, where N_MODELTASKS should be the ensemble size). One could also read the value from a configuration file.
Details on adapting a serial model
If the numerical model is not parallelized (i.e. serial), we need to add the parallelization for the ensemble. We follow here the approach used in the tutorial code tutorial/online_2D_serialmodel. The files mod_parallel_pdaf.F90, init_parallel_pdaf.F90, finalize_pdaf.F90 can be directly used. Note, that init_parallel_pdaf uses command-line parsing to read the number of model tasks (n_modeltasks) from the command line (specifying -dim_ens N_MODELTASKS, where N_MODELTASKS should be the ensemble size). One can replace this, e.g., by reading from a configuration file.
In the tutorial code, the parallelization is simply initialized by adding the line
CALL init_parallel_pdaf(1)
into the source code of the main program. This is done at the very beginning of the functional code part. This routine will call PDAF3_init_parallel to intialize and configure the parallelization.
The finalization of MPI is included in finalize_pdaf.F90 by a call to PDAF_finalize. The line
CALL finalize_pdaf()
should be inserted at the end of the main program.
With theses changes the model is ready to perform an ensemble simulation.
If the program is executed with these extensions using multiple model tasks, the issues discussed in 'Compiling the extended program' can occur. This one has to take care about which processes will perform output to the screen or to files.
Further Information
Communicators created for the data assimilation
MPI uses so-called 'communicators' to define groups of parallel processes. These groups can then conveniently exchange information. In order to provide the 2-level parallelism for PDAF, three communicators are initialized that define the processes that are involved in different tasks of the data assimilation system.
The required communicators are initialized in the call to PDAF3_init_parallel in the routine init_parallel_pdaf. They are called
COMM_model- defines the groups of processes that are involved in the model integrations (one group for each model task)COMM_filter- defines the group of processes that perform the filter analysis stepCOMM_couple- defines the groups of processes that are involved when data are transferred between the model and the filter. This is used to distribute and collect ensemble states (it is only used inside PDAF and not provided to the user code)
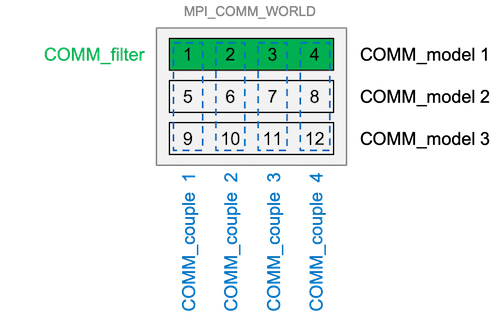
Figure 1: Example of a typical configuration of the communicators using a parallelized model. In this example we have 12 processes over all, which are distributed over 3 model tasks (COMM_model) so that 3 model states can be integrated at the same time. COMM_couple combines each set of 3 communicators of the different model tasks. The filter is executed using COMM_filter which uses the same processes of the first model tasks, i.e. COMM_model 1 (Figure credits: A. Corbin)
Arguments of init_parallel_pdaf
There are two variants of init_parallel_pdaf, one for a parallelized moden and one for a serial (or pure OpenMP) model.
For a parallelized model, the routine has the arguments
SUBROUTINE init_parallel_pdaf(screen, COMM_mymodel, rank_mymodel, size_mymodel)
with
screen: An integer defining whether information output is written to the screen (i.e. standard output). The following choices are available:- 0: quite mode - no information is displayed.
- 1: Display standard information about the configuration of the processes (recommended)
- 2: Display detailed information for debugging
COMM_mymodel: The model communicatorrank_mymodel: In integer giving therankof the process inCOMM_mymodel(usually initilized by a call to `MPI_Comm_rank)size_mymodel: In integer giving the number of processes inCOMM_mymodel(usually initilized by a call to `MPI_Comm_size)
For a serial model, the routine has one argument
SUBROUTINE init_parallel_pdaf(screen)
with
screen: An integer defining whether information output is written to the screen (i.e. standard output). The following choices are available:- 0: quite mode - no information is displayed.
- 1: Display standard information about the configuration of the processes (recommended)
- 2: Display detailed information for debugging
The arguments of PDAF3_init_parallel are described on the documentation page on PDAF3_init_parallel.
Compiling and testing the extended program
To compile the model with the adaption for modified parallelization, one needs to ensure that the additional files (init_parallel_pdaf.F90, mod_parallel_pdaf.F90, finalize_pdaf.F90) are included in the compilation.
If a serial model is used, one needs to adapt the compilation to compile for MPI-parallelization. Usually, this is done with a compiler wrapper like 'mpif90', which should be part of the OpenMP installation. If you compiled a PDAF tutorial case before, you can check the compile settings for the tutorial.
One can test the extension by running the compiled model (for the template in templates/online with mpirun -np 4 ./PDAF_online -dim_ens 4). With the default setting screen=1 in the call to init_parallel_pdaf, the standard output should include lines like
PDAF *** Initialize MPI communicators for assimilation with PDAF *** PDAF Pconf Process configuration: PDAF Pconf world assim model couple assimPE PDAF Pconf rank rank task rank task rank T/F PDAF Pconf ------------------------------------------------------------ PDAF Pconf 0 0 1 0 1 0 T PDAF Pconf 2 3 0 1 2 F PDAF Pconf 3 4 0 1 3 F PDAF Pconf 1 2 0 1 1 F
In this example only a single process will compute the filter analysis. There are 4 processes (with ranks 0 to 3) for 'world'. We have 4 model tasks (i.e. COMM_mymodel exists now 4 times), each using a single process, and one 'couple' comunicator with ranks 0 to 3. Thus, size_mycomm will be 1, and rank_mycomm will always be 0. (If one runs the exampe with mpirun -np 8 ./PDAF_online -dim_ens 4 one will get 8 lines instead of 4, size_mycomm will be 2 and rank_mycomm will show values 0 and 1.) Note that the couple communicator is only used internally by PDAF and is not accessible to the user code.
Using multiple model tasks can result in the following effects:
- The standard screen output of the model can by shown multiple times. For a serial model this is the case, since all processes can now write output. For a parallel model, this is due to the fact that often the process with
rank=0performs screen output. By splitting the communicatorCOMM_mymodel, each model task will have a process with rank 0. Here one can adapt the code to usemype_world==0frommod_parallel_pdaf. - Each model task might write file output. This can lead to the case that several processes try to generate the same file or try to write into the same file. In the extreme case this can result in a program crash. For this reason, it might be useful to restrict the file output to a single model task. This can be implemented using the variable
task_id, which is initialized byinit_parallel_pdafand holds the index of the model task ranging from 1 ton_modeltasks.- A convenient approach for the ensemble assimilation can be to run each model task in a separate directory. This needs an adapted setup. For example, for the tutorial in
tutorial/online_2D_parallelmodelwe can to the following:- in
tutorial/create sub-directoriesens1,ens2,ens3,ens4 - copy
online_2D_parallelmodel/model_pdafinto each of these directories - for ensemble size 4 and each model task using 2 processes, we can now run
mpirun -np 2 -wdir ens1 model_pdaf -dim_ens 4 : \ -np 2 -wdir ens2 model_pdaf -dim_ens 4 : \ -np 2 -wdir model_pdaf -dim_ens 4 : \ -np 2 -wdir model_pdaf -dim_ens 4
This approach has the advantage that each model writes files into a separate directory. This also allows to use the model's restart files, if it writes such.
- in
- One can also switch off the regular file output of the model completely. As each model task holds only a single member of the ensemble, this output might not be useful and might just slow down the program and lead to overly large use a disk space. In this case, the file output for the state estimate and perhaps all ensemble members should be done in the pre/poststep routine (
prepoststep_pdaf) of the assimilation system. This approach allows to run the full data assimilation in a single directory, but it can be combined with the use of separate directories.
- A convenient approach for the ensemble assimilation can be to run each model task in a separate directory. This needs an adapted setup. For example, for the tutorial in
If COMM_mymodel does not include all processes
If the parallelized model uses a communicator that is named different from MPI_COMM_WORLD (we use COMM_mymodel here), there can be two cases:
- The more typical case, described above is that
COMM_mymodelincludes all processes of the program. Often this is set asCOMM_mymodel = MPI_COMM_WORLDandCOMM_mymodelis introduced to have the option to define it differently. - In some models
COMM_mymodelis introduced to only represent a sub-set of the processes inMPI_COMM_WORLD. A use cases for this are coupled models, like atmosphere-ocean models. Here, the ocean might use a sub-set of processes and the program defines separate communicators for the ocean and atmosphere. There are also models that use a few of the processes to perform file operations with a so-called IO-server. In this case, the model runs only on nearly all processes andCOMM_mymodelwould be defined accordingly.
Case 2 is handled in PDAF V3.0 and later by PDAF because PDAF_init_parallel (or, for PDAF V3.0 the routine PDAF_set_parallel) is provided with the communicator COMM_mymodel and only reconfigures this communicator (thus, PDAF never uses MPI_COMM_WORLD.
Older implementations for PDAF 2.3.1 and before use a different approach. See the Page on adapting the parallelization for PDAF V2.x for more information.
